Welcome to Mrinversion documentation!¶
Deployment |


|
Build Status |

|
License |

|
Metrics |

|
About
The mrinversion python package is based on the statistical learning technique for
determining the underlying distribution of the magnetic resonance (NMR) parameters.
The library utilizes the mrsimulator package for generating solid-state NMR spectra and scikit-learn package for statistical learning.
Features
The mrinversion package includes
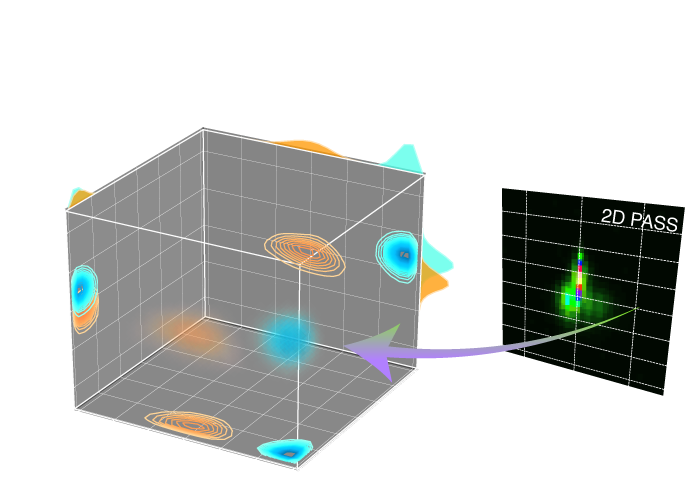
Spectral Inversion: Two-dimensional solid-state NMR spectrum of dilute spin-systems correlating the isotropic to anisotropic frequencies to a three-dimensional distribution of tensor parameters. Presently, we support the inversion of
Magic angle turning (MAT), Phase adjusted spinning sidebands (PASS), and similar spectra correlating the isotropic chemical shift resonances to pure anisotropic spinning sideband resonances into a three-dimensional distribution of nuclear shielding tensor parameters, \(\rho(\delta_\text{iso}, \zeta_\sigma, \eta_\sigma)\), where \(\delta_\text{iso}\) is the isotropic chemical shift, and \(\zeta_\sigma\) and \(\eta_\sigma\), are the shielding anisotropy and asymmetry parameters, respectively, defined using the Haeberlen convention.
Magic angle flipping (MAF) spectra correlating the isotropic chemical shift resonances to pure anisotropic resonances into a three-dimensional distribution of nuclear shielding tensor parameters, \(\rho(\delta_\text{iso}, \zeta_\sigma, \eta_\sigma)\), where \(\delta_\text{iso}\) is the isotropic chemical shift, and \(\zeta_\sigma\) and \(\eta_\sigma\), are the shielding anisotropy and asymmetry parameters, respectively, defined using the Haeberlen convention.
Relaxometry Inversion: Inversion of NMR relaxometry measurements to the distribution of relaxation parameters (T1, T2).
View our example gallery

Getting Started¶
Examples¶
Examples
Project details¶
Project details
How to cite¶
If you use this work in your publication, please cite the following.
Srivastava, D. J.; Grandinetti P. J., Statistical learning of NMR tensors from 2D isotropic/anisotropic correlation nuclear magnetic resonance spectra, J. Chem. Phys. 153, 134201 (2020). https://doi.org/10.1063/5.0023345.
Deepansh J. Srivastava, Maxwell Venetos, Philip J. Grandinetti, Shyam Dwaraknath, & Alexis McCarthy. (2021, May 26). mrsimulator: v0.6.0 (Version v0.6.0). Zenodo. http://doi.org/10.5281/zenodo.4814638
Additionally, if you use the CSDM data model, please consider citing
Srivastava DJ, Vosegaard T, Massiot D, Grandinetti PJ (2020) Core Scientific Dataset Model: A lightweight and portable model and file format for multi-dimensional scientific data. PLOS ONE 15(1): e0225953. https://doi.org/10.1371/journal.pone.0225953